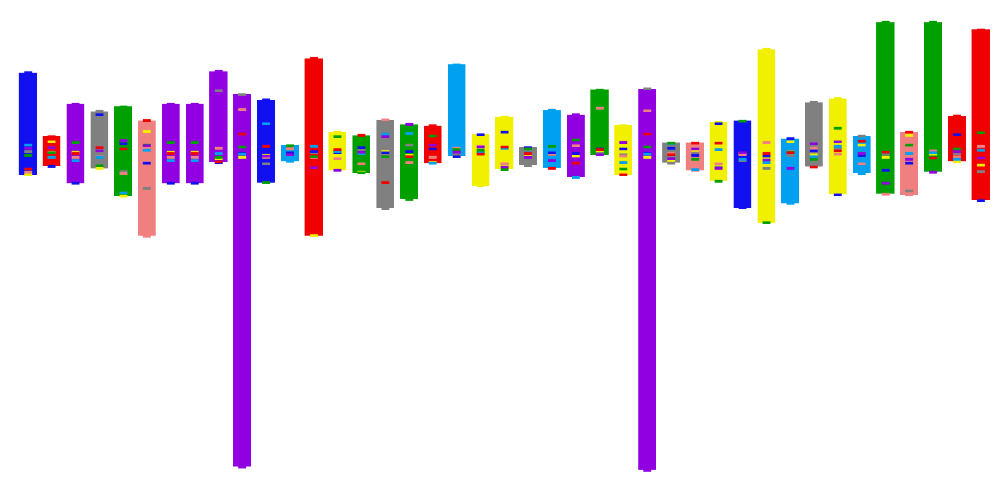
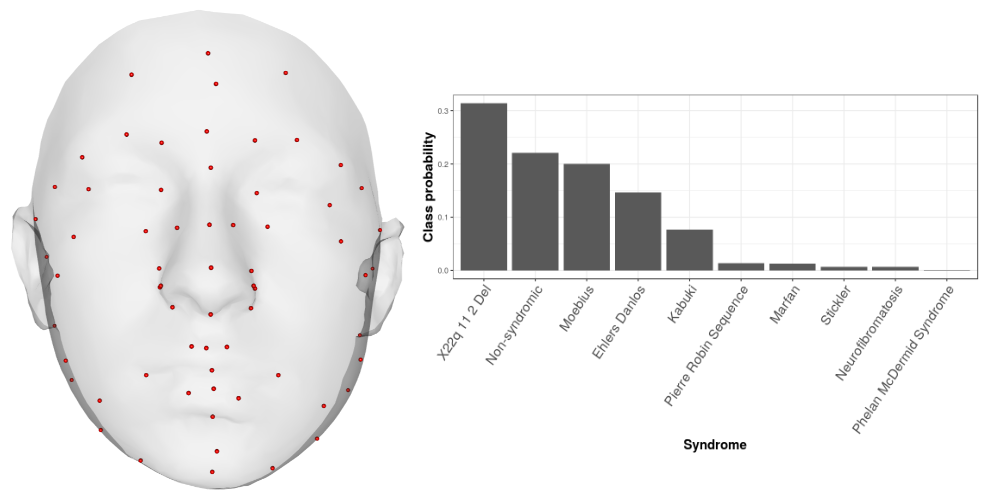
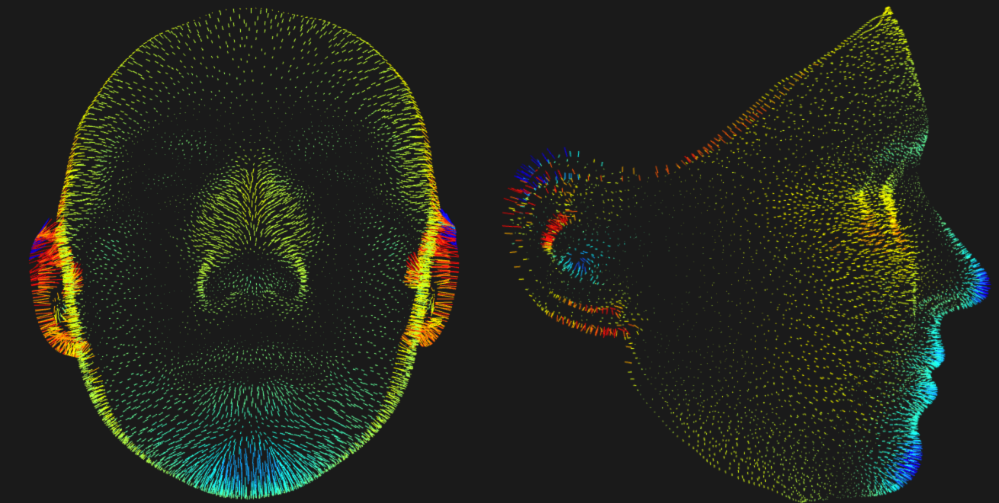
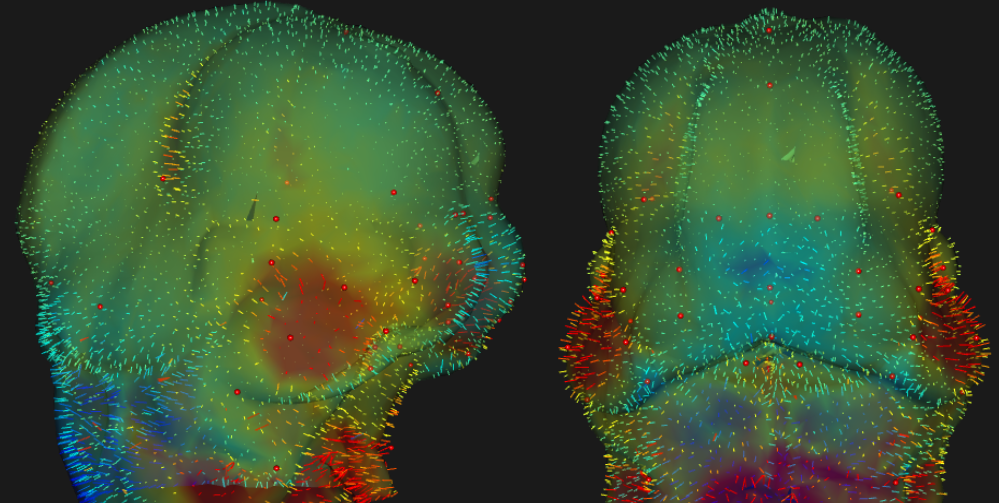
This app accompanies a paper all about the importance of development. We know that complex traits are structured by development, yet genomics seldom quantifies the importance of developmental pathways. That's what this app is all about.
Type some process you're interested in and you'll see what GO terms are listed under that process. From there, pick GO terms that you find interesting and we'll do the analysis right there and show you how alleles in our mouse sample covary with the shape of the face.
After you do all that, you can compare your pathway effect to any mutant in our database.